What we offer
Scientific Computing
We help you in performing scientific computing by applying the state-of-the-art techniques to:
- create and run data analysis workflows
- analyze and co-analyze your data and code
- make your code more reproducible
- modularize, optimize, parallelize and scale up your code
- use computing resources more efficiently (GPU, HPC cluster, cloud, ...).
We customize the service to the needs of research groups. Our past and running projects are in a diverse range of domains, e.g. natural language processing, bioinformatics, computational fluid dynamics, finite element analysis, etc.
Data Co-Analysis
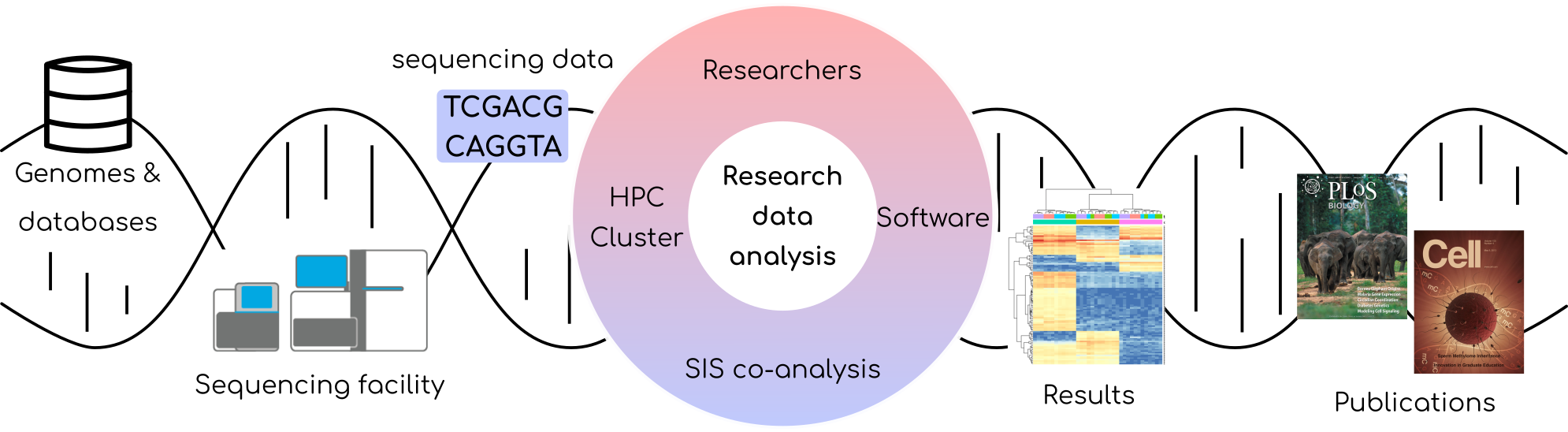
SIS is a member group of Swiss Institute of Bioinformatics (https://www.sib.swiss/bernd-rinn-group) thus provides bioinformatic support to the researchers who need to analyze their data in the areas of life sciences and biomedical research. In particular, we work in the co-analysis mode, where the user may decide on the type of support needed. Co-analysis projects range from complete outsourcing of computationally complex tasks to SIS, thorough analyzing the data together with PhD students and postdocs to on-demand consultancies and training for users who are advanced in bioinformatics. Our services are complementary to those delivered by core labs of ETH such as GFB in Basel (https://sis.id.ethz.ch/casestudies/case-study-genomics.html, https://bsse.ethz.ch/genomicsbasel) or FGCZ in Zurich (https://fgcz.ch/ ). In recent 4 years we have worked with research labs in over 50 co-analysis projects. Examples of their topic areas are:
- Parallelizing and scaling bioinformatic software on the computing cluster
- RNA-seq and genome sequencing data analysis
- Reproducible and automatized bioinformatic workflows, e.g. snakemake
- Non-standard genomic data analysis, e.g. alternative splicing
- Helping to deploy academic bioinformatic software or 3-rd party code
- Data mining in the outcome, e.g. search in sequences
- Code profiling and optimization (R, python, bash…)
Courses
We provide regular and customized trainings (see here).
Selected regular trainings:
- Research Data Management 2: Data Management and Analysis for reproducible research
- Parallel programming with MPI/OpenMP
- High-Performance Computing for genomic applications
